pacman::p_load(jsonlite, tidygraph, ggraph,
visNetwork, graphlayouts, ggforce,
skimr, tidytext, tidyverse)In-class Exercise 6b
6.0 VAST Challenge 2024 - MC3
6.1 Loading the necessary R packages
6.2 Data import
In the code chunk below, fromJSON() of jsonlite package is used to import MC3.json into the R environment
mc3_data <- fromJSON("data/MC3.json")In the code chunk below the class() function is used to confirm that the data is imported as a list
class(mc3_data)[1] "list"The output is called mc3_data. It is a large list R object.
6.3 Extracing edges
The code chunk below will be used to extract the links data.frame of mc3_data and saves it as a tibble data.frame called mc3_edges.
mc3_edges <-
as_tibble(mc3_data$links) %>%
distinct() %>% #to avoid duplicate records; source,target,type if they are the same will be treated as duplicates and kept as one
mutate(source = as.character(source),
target = as.character(target),
type = as.character(type)) %>%
group_by(source, target, type) %>%
summarise(weights = n()) %>% #weight - representation the number of linkages
filter(source!=target) %>%
ungroup()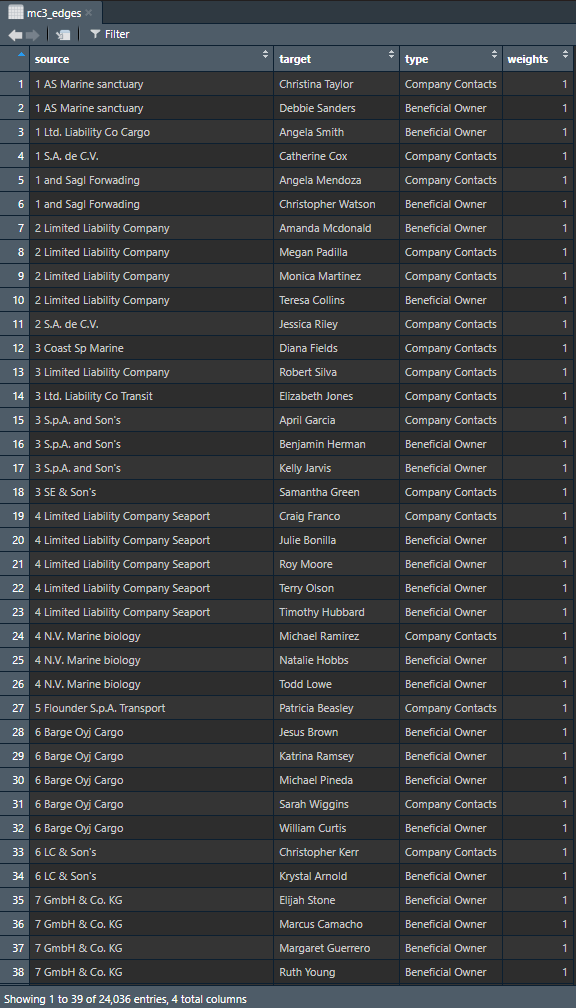
distinct()is used to ensure that there will be no duplicated records.mutate()andas.character()are used to convert the field data type from list to character.group_by()andsummarise()are used to count the number of unique links- the
filter(source!=target)is to ensure that no records are with similar source and target.
6.4 Extracing nodes
mc3_nodes <- as.tibble(mc3_data$nodes) %>% #extracting from the nodes data.frame
mutate(country = as.character(country),
id = as.character(id),
product_services = as.character(product_services),
revenue_omu = as.numeric(as.character(revenue_omu)), #as.numeric() since revenue should be processed as number to be able to do statistical analysis such as average
type = as.character(type)) %>%
select(id, country, type, revenue_omu, product_services)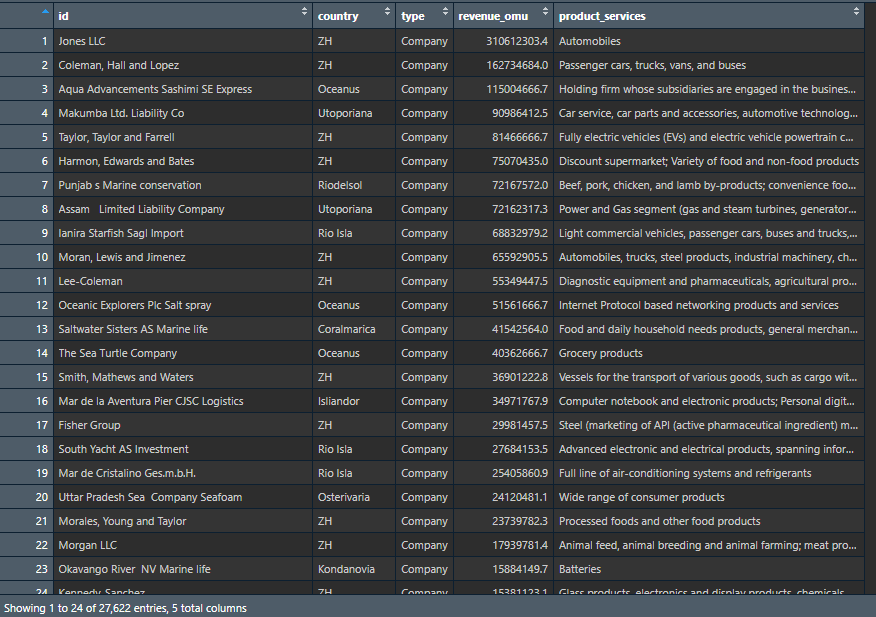
- to convert revenue_omu from list data type to numeric data type, we would have to convert the values into character first by using
as.character()followed byas.numeric()to convert the data into numeric data type. select()is used to re-organise the order of the fields.
6.5 Initial data exploration
Exploring the edges data frame
In the code chunk below, [skim()])(https://docs.ropensci.org/skimr/reference/skim.html) of skimr package is used to display the summary statistics of mc3_edges tibble data.frame.
skim(mc3_edges)| Name | mc3_edges |
| Number of rows | 24036 |
| Number of columns | 4 |
| _______________________ | |
| Column type frequency: | |
| character | 3 |
| numeric | 1 |
| ________________________ | |
| Group variables | None |
Variable type: character
| skim_variable | n_missing | complete_rate | min | max | empty | n_unique | whitespace |
|---|---|---|---|---|---|---|---|
| source | 0 | 1 | 6 | 700 | 0 | 12856 | 0 |
| target | 0 | 1 | 6 | 28 | 0 | 21265 | 0 |
| type | 0 | 1 | 16 | 16 | 0 | 2 | 0 |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| weights | 0 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 1 | ▁▁▇▁▁ |
The code function reveals that there is no missing value in the fields.
In the code chunk below, datatable() of DT package is used to display mc3_edges tibble data.frame as an interactive table on the html document.
DT::datatable(mc3_edges)ggplot(data = mc3_edges,
aes(x = type)) +
geom_bar() +
geom_text(aes(label = type), stat = "count", vjust = 5.5, colour = "white")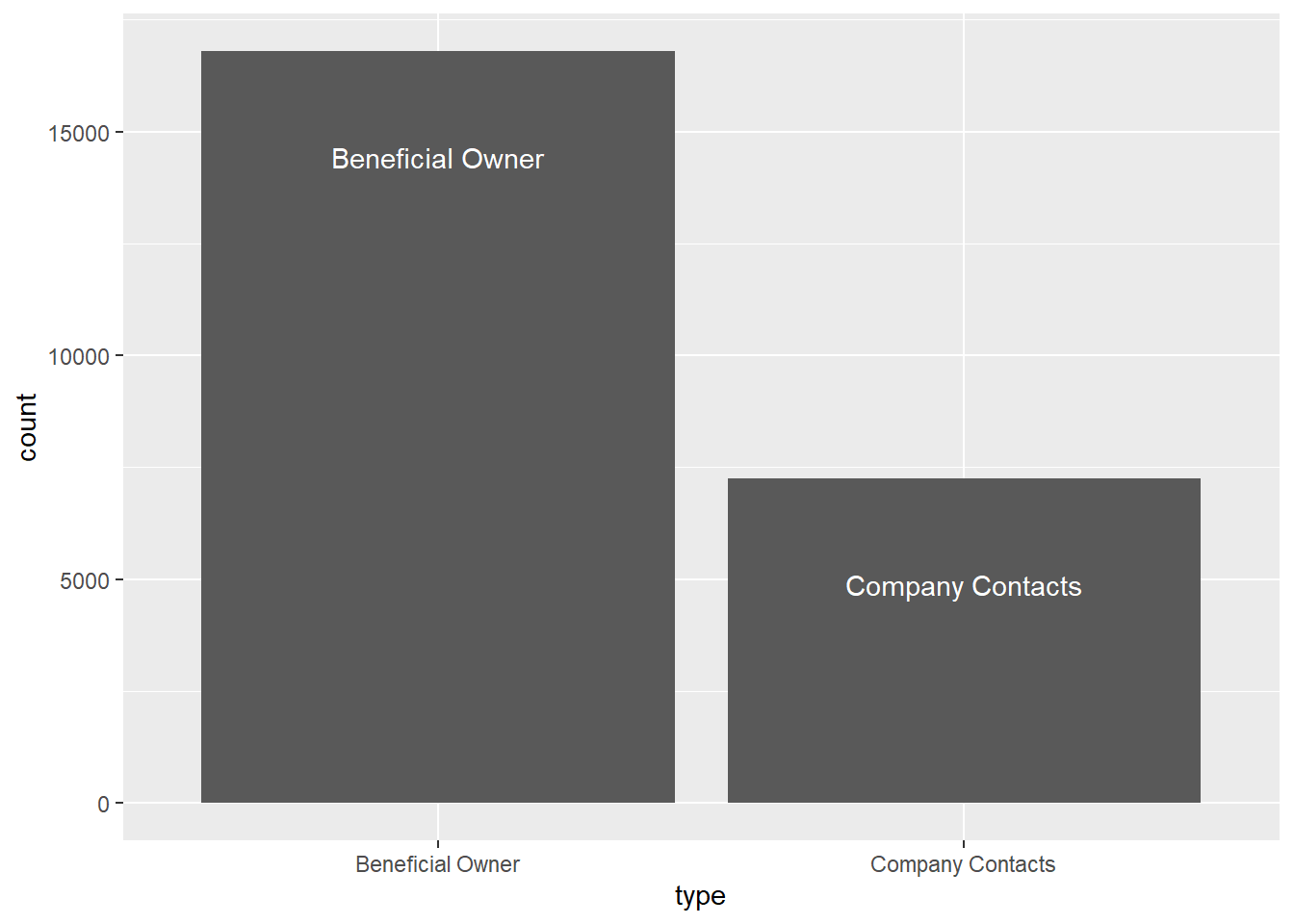
6.6 Building network model with tidygraph
6.6.1 Preparing the network graph
- since some data were trimmed away, the nodes and edges might not sync together
- from the code below, we pull out the name/ create a new node from the edges to represent source and target
- purpose is to update the nodes based on the new edges data in the R environment
- followed by combining id1 & id2 and using
left_join()to append them back to mc3_nodes
id1 <- mc3_edges %>%
select(source) %>%
rename(id = source)
id2 <- mc3_edges %>%
select(target) %>%
rename(id = target)
mc3_nodes1 <- rbind(id1, id2) %>%
distinct() %>%
left_join(mc3_nodes,
unmatched = 'drop')6.6.2 To construct the graph model
mc3_graph <- tbl_graph(nodes = mc3_nodes1,
edges = mc3_edges,
directed = FALSE) %>%
mutate(betweenness_centrality = # field name
centrality_betweenness(), # function - from tidygraph; nested function within the graph
closeness_centrality =
centrality_closeness())Building the graph to visualise
mc3_graph %>%
filter(betweenness_centrality >= 100000) %>% #`filter()` function used pull out nodes with betweenness centrality greater than input value.
ggraph(layout = "fr") +
geom_edge_link(aes(alpha=0.5)) +
geom_node_point(aes(
size = betweenness_centrality,
colors = "lightblue",
alpha = 0.5)) +
scale_size_continuous(range = c(1,10)) +
theme_graph()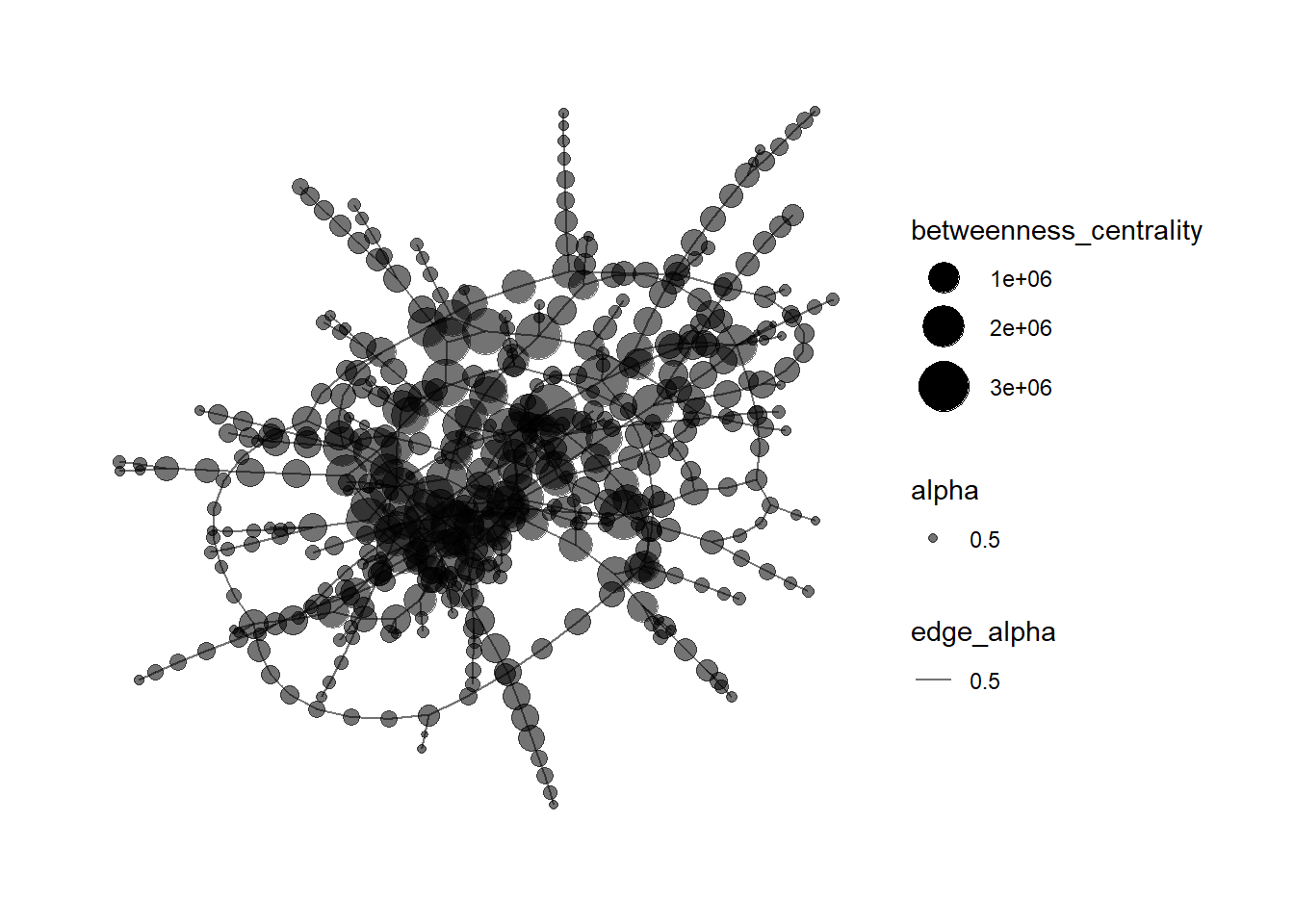
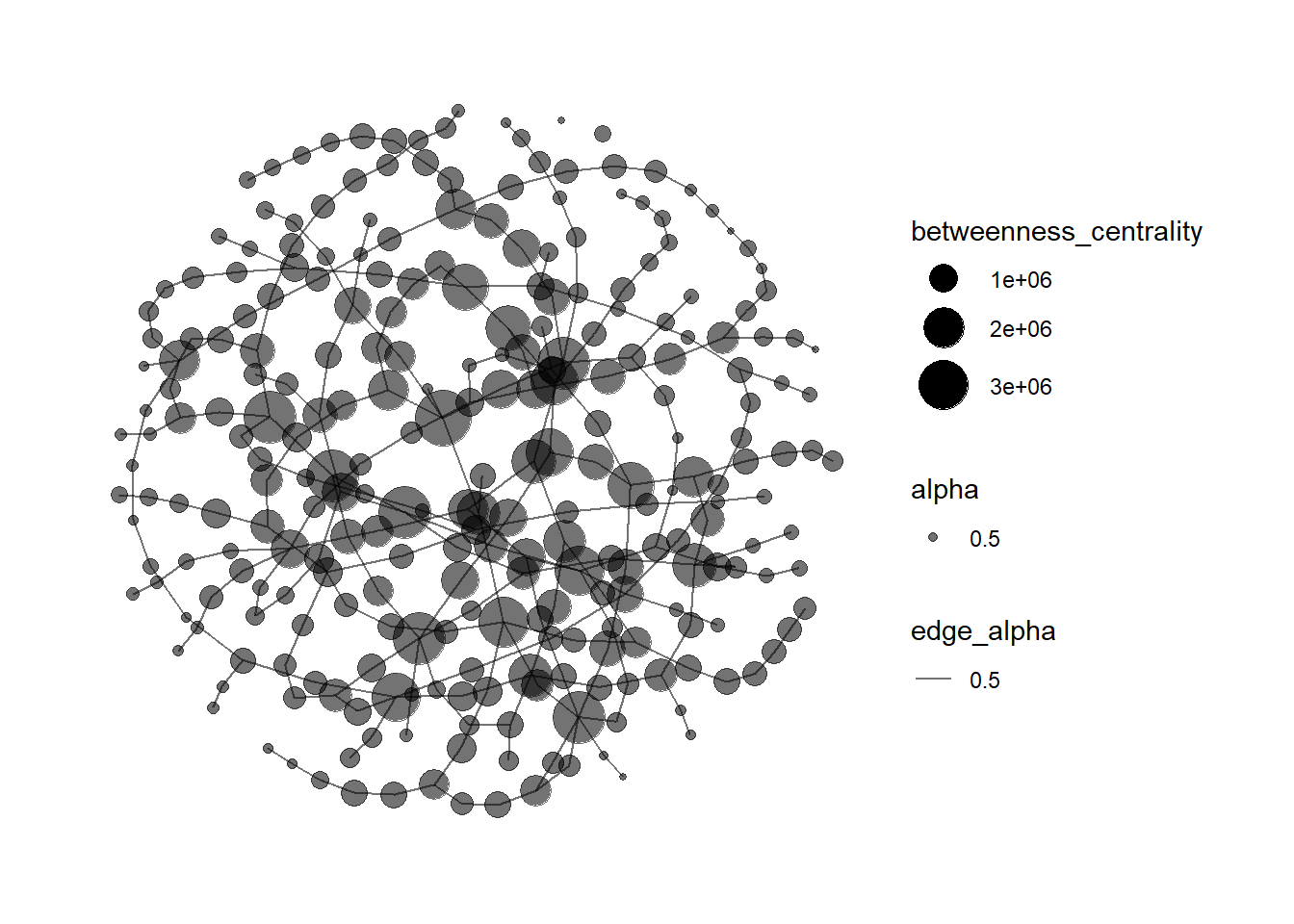
increased betweenness_centrality from 100000 to 300000;
mc3_graph %>%
filter(betweenness_centrality >= 300000) %>% #`filter()` function is used to pull out nodes with betweenness centrality greater than 300000 to avoid the graph being too cluttered.
ggraph(layout = "fr") +
geom_edge_link(aes(alpha=0.5)) +
geom_node_point(aes(
size = betweenness_centrality,
colors = "lightblue",
alpha = 0.5)) +
scale_size_continuous(range = c(1,10)) +
theme_graph()